More Information
Submitted: April 09, 2026 | Accepted: April 14, 2026 | Published: April 15, 2026
Citation: Mohamed SE, Elhussain KMK, Abdalla MHA. Genetic Association between the Endothelial Nitric Oxide Synthase Gene Polymorphism (T786C) and Nephropathy in Sudanese Sickle Cell Disease Patients in Khartoum State. J Hematol Clin Res. 2026; 10(1): 26-30. Available from:
https://dx.doi.org/10.29328/journal.jhcr.1001042
DOI: 10.29328/journal.jhcr.1001042
Copyright license: © 2026 Mohamed SE, et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Keywords: SCN; NO; eNOS; NOS3 (T786C)
Genetic Association between the Endothelial Nitric Oxide Synthase Gene Polymorphism (T786C) and Nephropathy in Sudanese Sickle Cell Disease Patients in Khartoum State
Samiah Edriss Mohamed1* , Khalid Mohamed Khalid Elhussain1,2 and Mahdi H. A. Abdalla2
, Khalid Mohamed Khalid Elhussain1,2 and Mahdi H. A. Abdalla2
1Omdurman Islamic University, Sudan
2Omdurman Ahlia University, Sudan
*Corresponding author: Samiah Edriss Mohamed, Omdurman Islamic University, Sudan, Email: [email protected]
Background: Sickle cell nephropathy (SCN) is one of the major cause of morbidity and mortality among patients with sickle cell disease (SCD), arising from chronic hemolysis, oxidative stress, and endothelial dysfunction. Variations in the endothelial nitric oxide synthase (eNOS) gene may influence nitric oxide (NO) bioavailability, potentially affecting renal susceptibility in SCD.
Aim: This study aimed to detect the association of eNOS gene polymorphism (T786C) with nephropathy among Sudanese patients with SCD.
Method: A case-control study was conducted between June 2021 and June 2022 at Soba University Hospital and Jafar Ibn Auf Specialized Hospital, Khartoum. 155 Participants included (65 patients with sickle cell nephropathy, two control groups 45 nephropathy patients without SCD, and 45 healthy controls). Samples were used for CBC and DNA extraction (RBCs, Hb, P CV, PLTs, WBCs, RBC count, and RBC indices): Was done using an automated hematology analyzer. Genomic DNA was extracted from blood samples and analyzed for T786C polymorphisms using polymerase chain reaction–restriction fragment length polymorphism (PCR-RFLP) techniques. Statistical assessment was carried out with a statistical package for the social sciences (SPSS), including Chi-square analysis and logistic regression.
Result: The NOS3 (T786C) genotype distributions differed significantly between SCD nephropathy cases and healthy controls (X² = 5.973, p = 0.050). The C allele of T786C was significantly more frequent among cases (p = 0.004). Logistic regression revealed a strong association of the T786C polymorphism with nephropathy under the dominant model, where carriers of at least one C allele (T/C+C/C) had increased risk compared to T/T carriers (OR = 8.96, 95%CI:1.11–72.14, p = 0.039). Genotype distribution of the normal control group predominantly exhibited the T/T genotype (97.8%), p = 0.050, with only1individual (2.2%) carryingtheT/Cgenotype, p = 0.050, and none with C/C, p = 0.050. The abnormal control group had 41 individuals (91.1%) with the T/T genotype, p = 0.050, 3 (6.7%) with T/C, p = 0.050, and 1 (2.2%) with C/C, p = 0.050. Hematological parameters showed no genotype- related variations.
Conclusion: eNOS T786C polymorphism appears to be associated with increased susceptibility to nephropathy among Sudanese SCD patients, suggesting a role for endothelial nitric oxide pathway dysfunction in the pathogenesis of sickle cell nephropathy.
The limitation faced is the sample size, which is sufficient to detect associations in dominant genetic models, but may lack the statistical power to capture subtle effects of less frequent genotypes.
Sickle cell nephropathy (SCN) is a serious, often under-recognized complication of sickle cell disease (SCD), contributing substantially to long-term morbidity and mortality. It encompasses a wide spectrum of renal abnormalities, ranging from asymptomatic tubular dysfunction and glomerular hyperfiltration to progressive proteinuria, chronic kidney disease, and ultimately end-stage renal disease. The onset of kidney involvement in SCD is often insidious, making early detection and risk stratification critical for improving outcomes [1].
The pathophysiology of SCN is complex, involving chronic hemolysis, oxidative stress, recurrent vaso-occlusion, and ischemia-reperfusion injury within the renal microvasculature. The renal medulla, characterized by low oxygen tension, hypertonicity, and acidosis, presents an environment that favors hemoglobin S polymerization and red cell sickling. These factors render the kidney, especially the medullary region, highly vulnerable to hypoxic damage and endothelial dysfunction [2,3]. One of the central mediators of endothelial homeostasis is nitric oxide (NO), a potent vasodilator synthesized by endothelial nitric oxide synthase (eNOS). NO plays a protective role by regulating vascular tone, inhibiting platelet aggregation, and reducing inflammation [4].
Nephropathy is a term used to describe a heterogeneous group of patients with Either microalbuminuria or varying degrees of proteinuria, with or without maternal hypertension or significant impairment in renal function. It is a broad medical term used to denote disease or damage of the kidney, which can eventually result in kidney failure. The primary and most obvious functions of the kidney are to excrete any waste products and regulate the water and acid-base balance of the body; therefore, loss of kidney function is a potentially fatal condition [5].
In SCD, impaired NO bioavailability is believed to contribute to vascular complications, including nephropathy.
The eNOS gene (NOS3), located on chromosome 7q36, encodes the enzyme responsible for constitutive NO production in endothelial cells. Polymorphisms in the eNOS gene have been shown to affect gene expression and enzymatic activity, potentially influencing NO levels and vascular function. Among the most widely studied variants are T786C, located in the promoter region, which is associated with reduced transcriptional activity, and G894T (Glu298Asp), a missense mutation that may impair enzyme stability and function [6,7].
While several studies have linked these polymorphisms to cardiovascular and renal diseases in various populations, data on their distribution and association with nephropathy in individuals with SCD, particularly in Sudan, remain limited. Given the high burden of both SCD and renal disease in this region, exploring the genetic determinants of nephropathy is essential for advancing risk prediction and personalized care [8].
Therefore, this study aimed to investigate the association of endothelial nitric oxide synthase gene polymorphisms (T786C) with nephropathy among Sudanese patients with sickle cell disease in Khartoum State.
This study was a case-control study conducted from June 2021 to June 2022 at two major Sudanese healthcare institutions: Soba University Hospital and Jafar Ibn Auf Specialized Hospital for Children, both located in Khartoum, Sudan. The study involved 155 participants, 65 sickle cell nephropathy cases, 36 (55.4%) male, and 29 (44.6%) were female, 45 non-Sickler nephropathy patients 28(62.2%) male and 17(37.8%) were female, and45 healthy control patients 23 (51.1%), 22 (48.9%) male and female, respectively. Exclusion criteria: patients with sickle cell nephropathy who had concurrent systemic illnesses and patients who declined genetic testing or sample provision. Venous blood (3 ml) was collected from each participant using EDTA tubes. Samples were used for CBC and DNA extraction. (Hb, PCV, PLTs, WBCs, RBC count, and RBC indices): Was done by using an automated hematology analyzer DIRUI(DIRUI BCC-3600). Genotyping of the T786C polymorphism in the NOS gene was performed using polymerase chain reaction-restriction fragment length polymorphism (PCR-RFLP) analysis, as described by Omneya -Moguib, et al. [9]. Genomic DNA was extracted from whole blood samples using the G-DEX™ IIb DNA extraction kit, following the manufacturer’s protocol. Statistical assessment was carried out with a statistical package for the social sciences (SPSS), including Chi-square analysis and logistic regression.
Ethical considerations
Ethical approval was obtained from the Ethical Committee of the Faculty of Medical Laboratory Sciences, Omdurman Islamic University, and the Khartoum State Ministry of Health. Written informed consent was obtained from all participants or their guardians. Permissions were granted by the administrative authorities of both hospitals. All collected data were handled with strict confidentiality and used solely for research purposes.
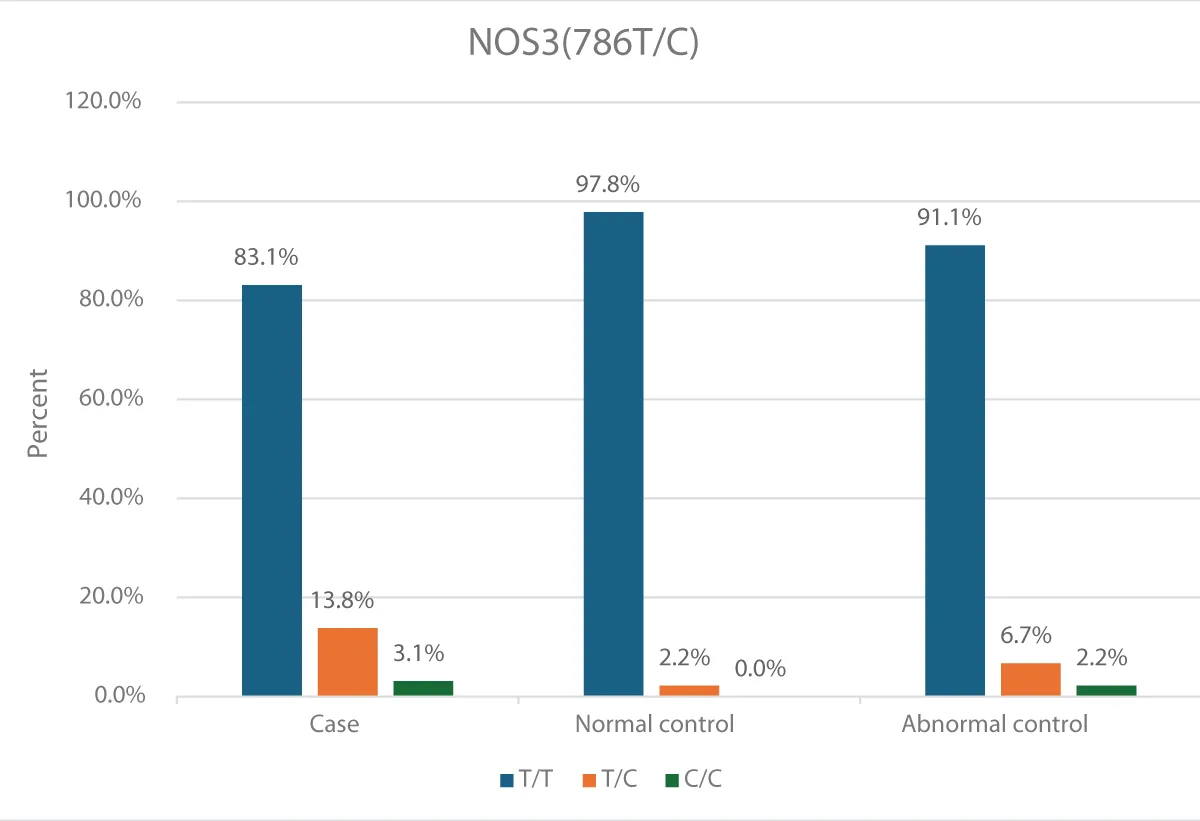
In this study, the distribution of the NOS3 (786T/C) polymorphism (Table 1, Figure 1), 54 patients (83.1%) in the case group carried the T/T genotype, 9 (13.8%) were heterozygous T/C, and 2 (3.1%) were homozygous for the C/C variant. The normal control group predominantly exhibited the T/T genotype (97.8%), withonly1individual (2.2%) carrying the T/C genotype and none with C/C. The abnormal control group had 41 individuals (91.1%) with the T/T genotype, 3 (6.7%) with T/C, and 1 (2.2%) with C/C.
| Table 1: Distribution of NOS3(786T/C) among study population | |||
| NOS3(786T/C) | Case | Normal control | Abnormal control |
| T/T | 54 (83.1%) | 44 (97.8%) | 41 (91.1%) |
| T/C | 9 (13.8%) | 1 (2.2%) | 3 (6.7%) |
| C/C | 2 (3.1%) | 0 (0.0%) | 1 (2.2%) |
| Total | 65 (100.0%) | 45 (100.0%) | 45 (100.0%) |
Figure 1: Distribution of NOS3(786T/C) among study population
Chi-square analysis in (Table2) revealed a significant difference was observed for the NOS3 (786T/C) genotype between cases and normal controls (X² = 5.982, p = 0.050), and for C allele carriers specifically (X² = 8.333, p = 0.004). However, in Table 3, no statistically significant
| : Distribution of NOS3(786T/C) polymorphisms between Sickler patients and normal controls | ||||
| Genotype/Allele | Sickler cases(n= 65) | Normal controls(n=45) | Chi-square(X²) | p - value |
| NOS3(786T/C) | ||||
| T/T | 54 (83.1%) | 44 (97.8%) | 5.982 | 0.050* |
| T/C | 9 (13.8%) | 1 (2.2%) | ||
| C/C | 2 (3.1%) | 0 (0.0%) | ||
| Callelecarriers(T/C+C/C) | 11 (16.9%) | 1 (2.2%) | 8.333 | 0.004* |
| Table 3: Distribution of NOS3(786T/C) polymorphisms between Sickler patients and abnormal controls | ||||
| Genotype/Allele | Sickler Cases(n=65) | Abnormal Controls(n=45) | Chi-square(X²) | p - value |
| NOS3(786T/C) | ||||
| T/T | 54 (83.1%) | 41 (91.1%) | 1.526 | 0.466 |
| T/C | 9 (13.8%) | 3 (6.7%) | ||
| C/C | 2 (3.1%) | 1 (2.2%) | ||
| C allele carriers(T/C+C/C) | 11 (16.9%) | 4 (8.9%) | 3.267 | 0.071 |
Significant differences were found when comparing cases with abnormal controls for either polymorphism (all p > 0.05).
Table 4 summarizes the logistic regression analysis under various genetic models. In the dominant genetic model analysis of NOS3 (786T/C) between cases and normal controls, individuals carrying at least one C allele (T/C+C/C) showed a significant association with nephropathy compared to T/T carriers (OR = 8.963, 95% CI: 1.114 – 72.136, p = 0.039).
| Table 4: Genetic Model Analysis of NOS3(786T/C) polymorphisms between cases and control groups | |||||
| Polymorphism | Genetic model (Cases vs.) | Comparison | p - value | Odds ratio (OR) | 95% C.I |
| NOS3(786T/C) | Dominant (vs. Normal Control) | T/C+C/C vs. T/T | 0.039* | 8.963 | 1.114-72.136 |
| NOS3(786T/C) | Codominant (vs. Abnormal Control) | T/C vs. T/T | 0.737 | 1.519 | 0.133-17.327 |
| NOS3(786T/C) | Codominant(vs. Abnormal Control) | C/Cvs.T/T | 0.771 | 0.667 | 0.043-10.253 |
For the codominant genetic model analysis of NOS3(786T/C) between cases and abnormal controls did not reveal significant associations for either T/C (OR = 1.519, 95%CI:0.133 – 17.327, p = 0.737) or C/C (OR = 0.667, 95% CI: 0.043 – 10.253, p = 0.771) genotypes compared to T/T. When comparing cases to abnormal normal controls under the dominant model of NOS3(786T/C), no significant association was observed (OR = 0.479, 95% CI: 0.142–1.613, p = 0.235). Similarly, in the recessive genetic model of NOS3(786T/C), no significant association was detected between cases and abnormal normal controls (OR = 1.397, 95% CI: 0.123 – 15.885, p = 0.788). Table 5 show no significant difference was observed for the NOS3(786T/C) genotype between Hematological parameters and NOS3 (786T/C) genotype.
| : Comparison of hematological parameter according of eNOS (786T/C) polymorphisms | ||||||||
| eNOS (786T/C) | RBCS | Hb | PCV | MCV | MCH | MCHC | TWBCs | Platelet |
| T/T | 2.5 ± 0.6 | 6.8 ± 1.6 | 22.2 ± 4.3 | 80.9 ± 7.3 | 26.6 ± 2.3 | 32.0 ± 2.6 | 11.9 ± 6.4 | 217.1 ± 85.3 |
| T/C | 2.5 ± 0.5 | 6.7 ± 1.4 | 22.8 ± 4.5 | 82.5 ± 6.5 | 26.9 ± 2.1 | 34.7 ± 3.7 | 11.4 ± 3.9 | 193.1 ± 22.8 |
| C/C | 1.9 ± 0.7 | 5.8 ± 2.4 | 17.8 ± 5.2 | 85.3 ± 0.9 | 28.8 ± 0.5 | 32.1 ± 6.8 | 10.3 ± 7.1 | 320.5 ± 127.9 |
| P. value | 0.506 | 0.695 | 0.344 | 0.609 | 0.412 | 0.047* | 0.908 | 0.141 |
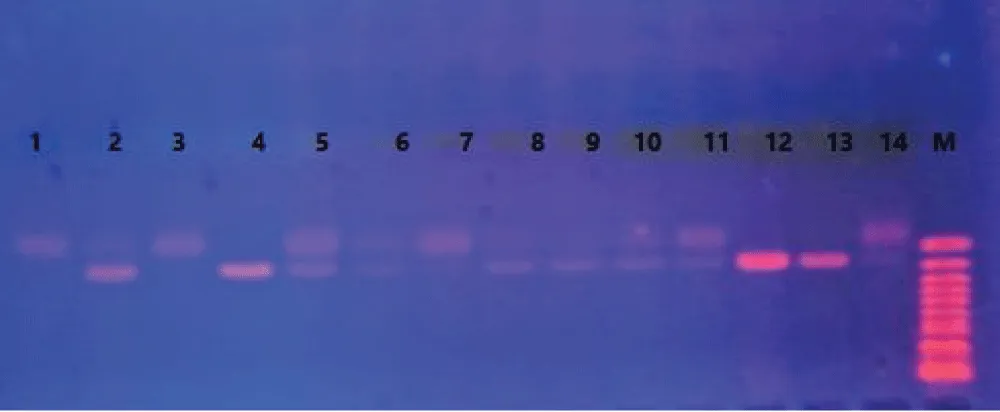
Figure 2 shown PCR products of NOS3 gene (-786T/C) (rs2070744 ) in 1.5% agarose gel in1XTBE ,stained with 0.5µg /ml Ethidium bromide (digested with MSP1 enzyme) yield fragments of 140 and 40 could not be reseloved in the gel(only 140 band was seen) Lane 1, 3 and 6 and (mutant 90 bp, 50 and 40 could not be seen in the gel) C/C Lane4, 12 and 13 wild type (140 bp) T/T Lane 2, 5, 6, 8,9,10,11 and 14 heterozygous(140,90)T/C.
Figure 2: PCR products of NOS3 gene (-786T/C) (rs2070744).
Notably, not all polymorphisms demonstrated sufficient genotype variation across all study groups to permit full application of all genetic models. In some comparisons, certain genotypes (such as C/C or T/T) were either absent or extremely rare, limiting the feasibility of codominant and recessive analyses. These limitations were considered in the model selection process, and only statistically valid comparisons based on available genotype distributions were included in the final analysis.
This study investigated the distribution and association of NOS3 (T786C) polymorphisms among patients with sickle cell nephropathy in comparison to normal healthy controls and nephropathy patients without sickle cell disease (SCD).We found that the frequency of the C allele and C-containing genotypes of the NOS3 T786C polymorphism was significantly higher in the SCD nephropathy group compared to normal healthy controls, indicating that this variant is associated with an increased risk of nephropathy in sickle cell patients.
Our findings align with the review by Medina, et al. [10], which highlighted that NOS3 polymorphisms, including G894T, T786C, and intron 4 VNTR variants, accelerate kidney function decline through endothelial dysfunction and oxidative stress pathways. Importantly, Medina, et al. emphasized that the association of these variants with chronic kidney disease (CKD) exhibits ethnic and population variability, consistent with the variable genotype distributions observed in our Sudanese cohort. Similar observations were reported by Tanus- Santos, et al. [11], who described significant differences in NOS3 polymorphism frequencies across ethnicities, under scoring the importance of population-specific genetic investigations.
Although direct studiesonNOS3 polymorphisms and sickle cell nephropathy are limited, the pathophysiology of SCD nephropathy shares mechanisms with other nephropathies, particularly endothelial dysfunction. Naik and Derebail [12] reviewed the spectrum of sickle hemoglobin-related nephropathy and highlighted impaired NO signaling, where eNOS/NOS3 plays a central role, as a key contributor to glomerular injury and progressive CKD in SCD patients.
Our study adds genetic evidence supporting the role of NOS3 polymorphisms, especially T786C, in modulating nephropathy risk among SCD patients. Nishank, et al. [13] reported similar findings in Indian SCD patients, where the T786C polymorphism was significantly associated with SCD complications. Conversely, Thakur, et al. (2014) in Malian SCD patients and Navarro, et al. [8,14] in African-American populations reported no significant differences inT786Callele and genotype distributions between patients and controls, reflecting the ethnic and geographical variability in genetic susceptibility, which is consistent with our observations.
Notably, the absence of significant differences in hematological indices (RBCs,Hb,PCV, TWBCs, and platelets) across NOS3 genotypes in our cohort indicates that these polymorphisms likely exert their effect via endothelial and vascular pathways rather than through direct hematological alterations.
Regarding the abnormal control group(patients with nephropathy but without SCD), the lack of significant genotype distribution differences compared to SCD nephropathy cases suggests that NOS3 polymorphisms may represent a general genetic risk factor for nephropathy, irrespective of underlying etiology. This observation supports the notion that endothelial dysfunction is a converging pathogenic mechanism across various nephropathies.
Studies investigating NOS3 polymorphisms in nephropathy have primarily focused on diabetic populations. Armenis et al. [15] demonstrated a significant association between NOS3 polymorphisms and diabetic nephropathy in Greek patients, while Dellameaetal [16] reported similar findings in a systematic review and meta-analysis. Although these studies pertain to diabetic nephropathy, they further highlight the universal impact of NOS3 genetic variability on renal disease susceptibility across different clinical settings. The lack of available studies specifically exploring NOS3 polymorphisms in sickle cell nephropathy underscores the novelty of our findings and the need for further research.
In alignment with our genetic findings, Chenou, et al. [17] conducted a study on Brazilian sickle cell anemia patients, assessing eNOS polymorphisms (T786C, G894T, and VNTR intron 4) alongside markers of hemolysis, inflammation, and endothelial dysfunction. The study found significantly higher frequencies of the −786C allele and C/C genotype among SCD patients compared to healthy controls, although no significant correlation was observed with inflammatory or hemolytic markers [17]. Their findings are in agreement with Nishank, et al. [13], but contrary to Thakuretal [8] and Navarro, et al. [14], who found no association. Additionally, Chenou, et al. [17] highlighted the complex interplay between eNOS polymorphisms and endothelial dysfunction markers, suggesting that these genetic variants.
May contribute to phenotypic variability and disease progression in SCD through endothelial pathways.
Strengths and limitations
A major strength of this study is
- Well-characterized case-control design,
- Incorporating both healthy and nephropathy controls, enabling a more precise assessment of genotype-disease associations.
- The inclusion of Sudanese Sicklers also provides valuable insights from a region with limited genetic epidemiological data.
- The sample size, while sufficient to detect associations in dominant genetic models, may lack the statistical power to capture subtle effects of less frequent genotypes.
- Cretin genotypes were either absent or extremely rare, limiting the feasibility of co dominant and recessive analyses. These limitations were considered in the model selection process, and only statistically valid comparisons based on available genotype distributions were included in the final analysis.
This study provides novel evidence that the NOS3 T786C polymorphism is significantly associated with an increased risk of nephropathy in Sudanese patients with sickle cell disease, highlighting the important role of endothelial nitric oxide synthase in the pathogenesis of sickle cell nephropathy. The findings also suggest that NOS3 genetic variants may represent common risk factors for nephropathy beyond sickle cell disease, as indicated by the similar genotype distributions observed in nephropathy patients without SCD.
Implications and future directions
The present study implicates the NOS3 T786C polymorphism as a potential genetic risk marker for nephropathy in SCD patients. Identification of individuals carrying this risk allele could aid in risk stratification and inform early intervention strategies aimed at preserving renal function. Future research should focus on larger, multicenter cohorts to validate these findings, including functional studies evaluating endothelial function and NO levels in relation to NOS3 genotypes. Moreover, integrating additional genetic modifiers, such as APOL1 variants, and conducting haplotype analyses may provide a more comprehensive understanding of genetic susceptibility to nephropathy in SCD.
The authors would like to thank Transasia Biomedicals for supporting with the new Hematology analyzer Erba H7100 and its reagent for study purpose along with indispensable support from their Product management team including Deepak Valand & Trushi Sonavaria along with Rahat Shaikh who supported in installation and processing the samples for the study.
- Baddam S, Rout P. Sickle cell nephropathy. Natl Libr Med. Jan 2025. Available from: https://www.ncbi.nlm.nih.gov/books/NBK526017/
- Pham PT. Renal abnormalities in sickle cell disease. Kidney Int. 2000;57:1-8. Available from: https://doi.org/10.1046/j.1523-1755.2000.00806.x
- Ataga KI. Renal abnormalities in sickle cell disease. Am J Hematol. 2000;63:205-11. Available from: https://doi.org/10.1002/(sici)1096-8652(200004)63:4%3C205::aid-ajh8%3E3.0.co;2-8
- Dierk H, Ernesto S. Endothelial dysfunction. Natl Libr Med. 2004;15(8):1983-92. Available from: https://doi.org/10.1097/01.asn.0000132474.50966.da
- Mestrovic T. What is nephropathy. News Med Life Sci. Mar 2023. Available from: https://www.news-medical.net/health/What-is-Nephropathy.aspx
- Wang Y, Marsden PA. Nitric oxide synthases: gene structure and regulation. Natl Libr Med. 1995;34:71-90. Available from: https://doi.org/10.1093/eurheartj/ehr304
- Moguib O, Raslan H, Abdel Rasheed I, Effat L, Mohamed N, Thakur TJ, et al. Endothelin-1 but not endothelial nitric oxide synthase gene polymorphism is associated with sickle cell disease in Africa. Natl Libr Med. 2014;25(8):119-26.
- Thakur TJ, Guindo A, Cullifer LR, Li Y, Imumorin IG, Diallo DA, Thomas BN. Endothelin-1 but not endothelial nitric oxide synthase gene polymorphism is associated with sickle cell disease in Africa. Natl Libr Med. 2014;25(8):119-26. Available from: https://doi.org/10.4137/grsb.s14836
- Cruz-Gonzalez I, Corral E, Sanchez-Ledesma M, Sanchez-Rodriguez A, Martin-Luengo C, Gonzalez-Sarmiento R. Association between T-786C NOS3 polymorphism and resistant hypertension: prospective cohort study. BMC Cardiovasc Disord. 2009;9(1):35. Available from: https://doi.org/10.1186/1471-2261-9-35
- Medina AM, Zubero EE, Jiménez MAA, Barragan SAA, García CAL, Ramos JJG, et al. NOS3 polymorphisms and chronic kidney disease. J Bras Nefrol. 2018;40(3):273-7. Available from: https://doi.org/10.1590/2175-8239-JBN-3824
- Tanus-Santos JE, Desai M, Flockhart DA. Effects of ethnicity on the distribution of clinically relevant endothelial nitric oxide variants. Pharmacogenet Genomics. 2001;11(8):719-25. Available from: https://doi.org/10.1097/00008571-200111000-00011
- Naik RP, Derebail VK. The spectrum of sickle hemoglobin-related nephropathy: from sickle cell disease to sickle trait. Expert Rev Hematol. 2017;10(12):1087-94. https://doi.org/10.1080/17474086.2017.1395279
- Nishank SS, Singh MP, Yadav R, Gupta RB, Gadge VS, Gwal A. Endothelial nitric oxide synthase gene polymorphism is associated with sickle cell disease patients in India. J Hum Genet. 2013;58(12):775-9. https://doi.org/10.1038/jhg.2013.99
- Navarro KG, Agyingi SE, Nwabuobi CK, Thomas BN. Polymorphism of the endothelin-1 gene (rs5370) is a potential contributor to sickle cell disease pathophysiology. Genes Dis. 2016;3(4):294-8. https://doi.org/10.1016/j.gendis.2016.09.002
- Armenis I, Kalotychou V, Tzanetea R, Konstantopoulos K, Rombos I. The effect of endothelial nitric oxide synthase G894T and T786C polymorphisms on hypoxia-inducible factor-1 alpha expression in sickle cell disease. Nitric Oxide. 2021;111:31-6. Available from: https://doi.org/10.1016/j.niox.2021.03.004
- Dellamea BS, Pinto LCF, Leitão CB, et al. Endothelial nitric oxide synthase gene polymorphisms and risk of diabetic nephropathy: a systematic review and meta-analysis. BMC Med Genet. 2014;15:9. Available from: https://doi.org/10.1186/1471-2350-15-9
- Chenou F, Albuquerque DM, Leonardo DP, Domingos IF, Bezerra MAC, Araújo AS, et al. Endothelial nitric oxide synthase (eNOS) gene polymorphisms and markers of hemolysis, inflammation and endothelial dysfunction in Brazilian sickle cell anemia patients. Biochem Genet. 2020;58(4):580-94. Available from: https://doi.org/10.1007/s10528-020-09959-w